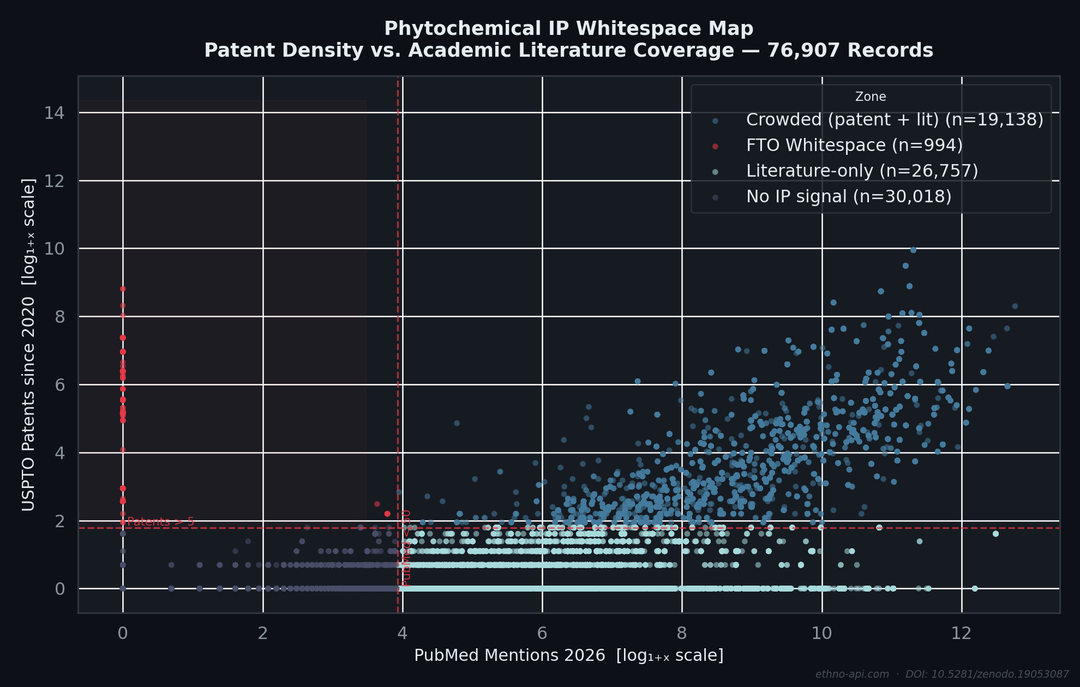
Each point represents a phytochemical from the USDA’s Dr. Duke database, plotted against patents filed with the USPTO since 2020 (y-axis) and the citation frequency in PubMed (x-axis). Both axes are logarithmically scaled.
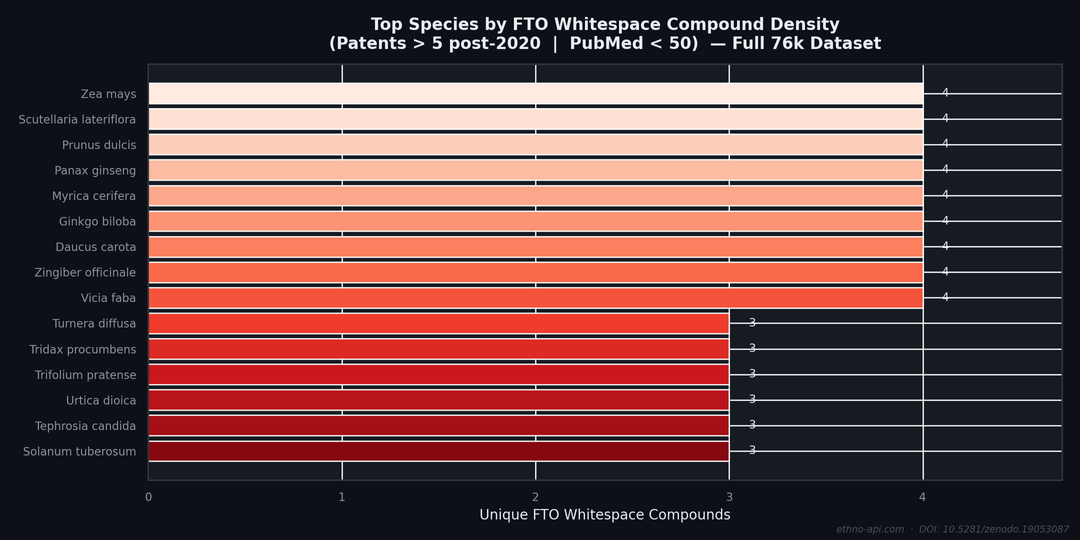
The red area: high patent density, low scientific literature—this is what IP analysts refer to as FTOwhitespace: commercial activities that have not yet resulted in peer-reviewed scientific publications. In a sample of 400 records, the query returns compounds with more than 5 patents and fewer than 50 citations in PubMed.
Created from a flat dataset of 76,000 records that combines USDA ethnobotanical records with PubMed, ClinicalTrials.gov, ChEMBL bioactivity data, and PatentsView. The complete pipeline is available in the GitHub repository, including the DuckDB query and the ChromaDB RAG embedding.
github.com/wirthal1990-tech/USDA-Phytochemical-Database-JSON
by DoubleReception2962
1 Comment
**Source:** USDA Dr. Duke’s Phytochemical and Ethnobotanical Databases (public domain) — denormalized and enriched with:
– PubMed citation counts via NCBI E-utilities
– [ClinicalTrials.gov](http://ClinicalTrials.gov) study counts (API v2)
– ChEMBL bioactivity measurements (with PubChem InChIKey fallback)
– USPTO patent counts via PatentsView (post-2020)
Full dataset: 76,907 records across 24,746 unique compounds and 2,313 plant species.
DOI: 10.5281/zenodo.19053087
**Tool:** Python (matplotlib + seaborn), DuckDB for the FTO whitespace query. Both axes are log₁₊ₓ scaled to handle the heavy right-skew in citation counts.
**Code + methodology:**
[github.com/wirthal1990-tech/USDA-Phytochemical-Database-JSON](http://github.com/wirthal1990-tech/USDA-Phytochemical-Database-JSON)
The full pipeline including the DuckDB query used to classify compounds into the four zones (FTO Whitespace / Crowded / Literature-only / No IP signal) is documented in [METHODOLOGY.md](http://METHODOLOGY.md) in the repo.